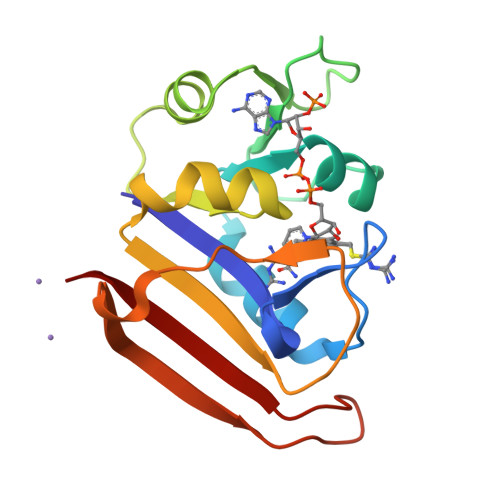
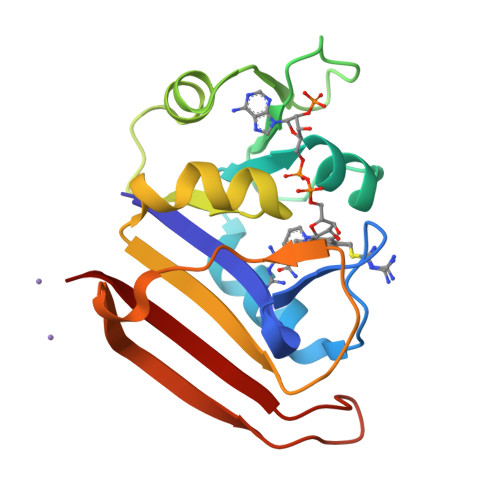
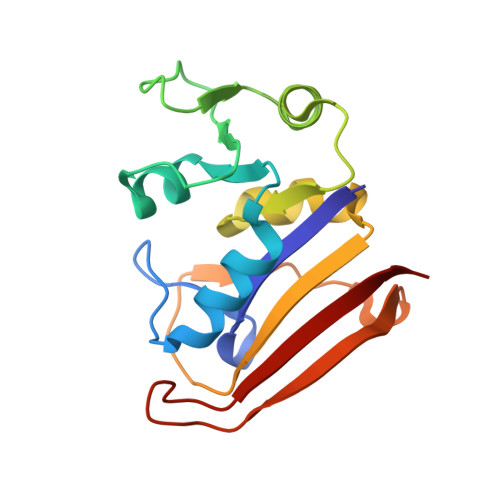
A 2.13 A Structure of E. coli Dihydrofolate Reductase Bound to a Novel Competitive Inhibitor Reveals a New Binding Surface Involving the M20 Loop Region
Summerfield, R.L., Daigle, D.M., Mayer, S., Mallik, D., Hughes, D.W., Jackson, S.G., Sulek, M., Organ, M.G., Brown, E.D., Junop, M.S.(2006) J Med Chem 49: 6977-6986
- PubMed: 17125251
- DOI: https://doi.org/10.1021/jm060570v
- Primary Citation of Related Structures:
2ANO, 2ANQ - PubMed Abstract:
Dihydrofolate reductase (DHFR) is a vital metabolic enzyme and thus a clinically prominent target in the design of antimetabolites. In this work, we identify 1,4-bis-{[N-(1-imino-1-guanidino-methyl)]sulfanylmethyl}-3,6-dimethyl-benzene (compound 1) as the correct structure of the previously reported DHFR inhibitor 1,4-bis-{(iminothioureidomethyl)aminomethyl}-3,6-dimethyl-benzene (compound 2). The fact that compound 1 has an uncharacteristic structure for DHFR inhibitors, and an affinity (KI of 11.5 nM) comparable to potent inhibitors such as methotrexate and trimethoprim, made this inhibitor of interest for further analysis. We have conducted a characterization of the primary interactions of compound 1 and DHFR using a combination of X-ray structure and SAR analysis. The crystal structure of E. coli DHFR in complex with compound 1 and NADPH reveals that one portion of this inhibitor exploits a unique binding surface, the M20 loop. The importance of this interface was further confirmed by SAR analysis and additional structural characterization.
Organizational Affiliation:
Atlantic Veterinary College, University of Prince Edward Island, 550 University Avenue, Charlottetown, Prince Edward Island C1A 4P3, Canada.